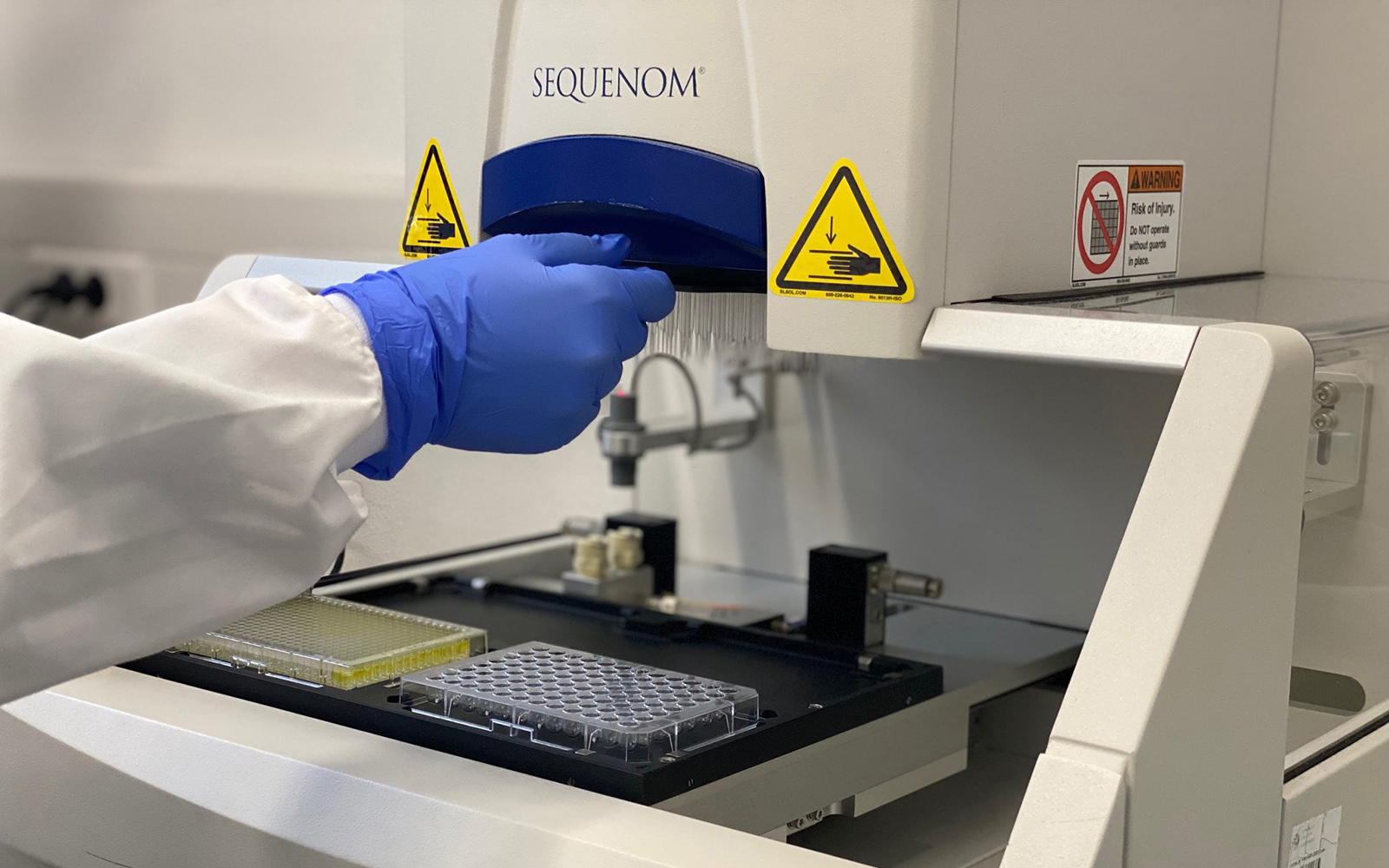
Genomics
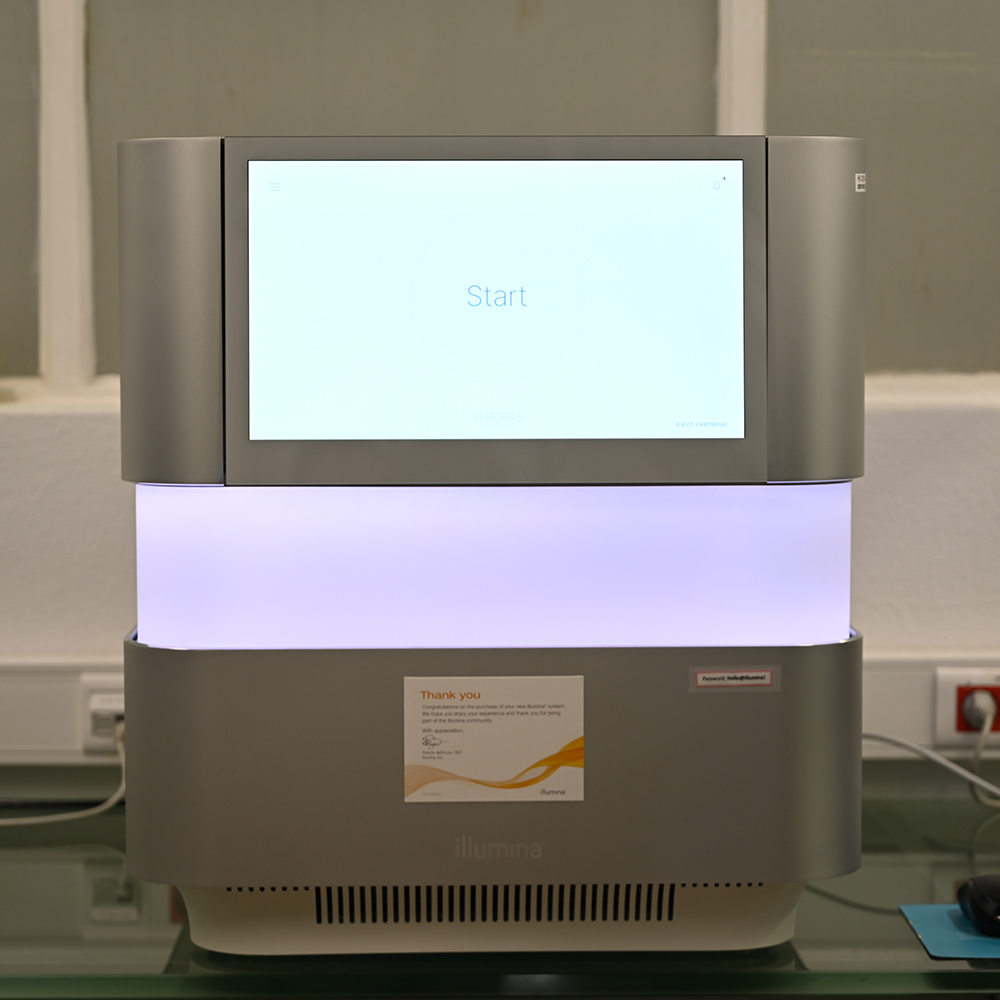
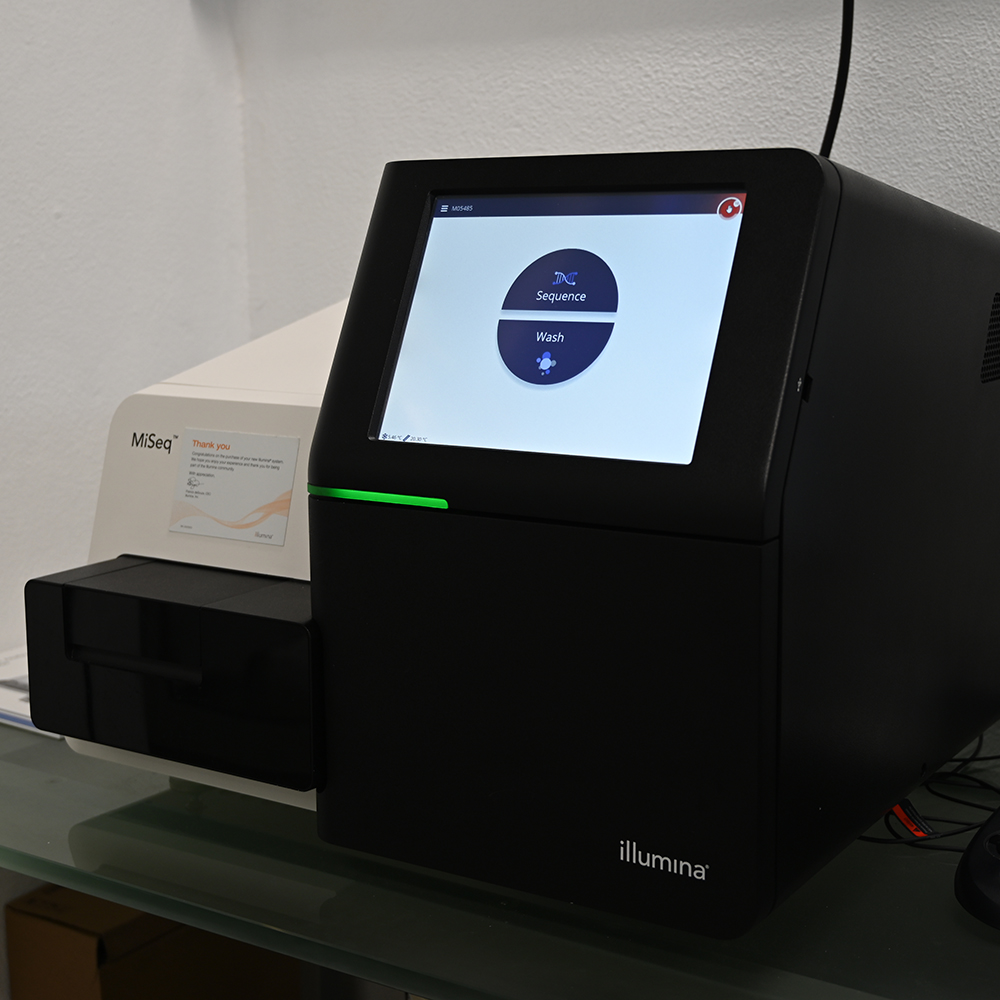
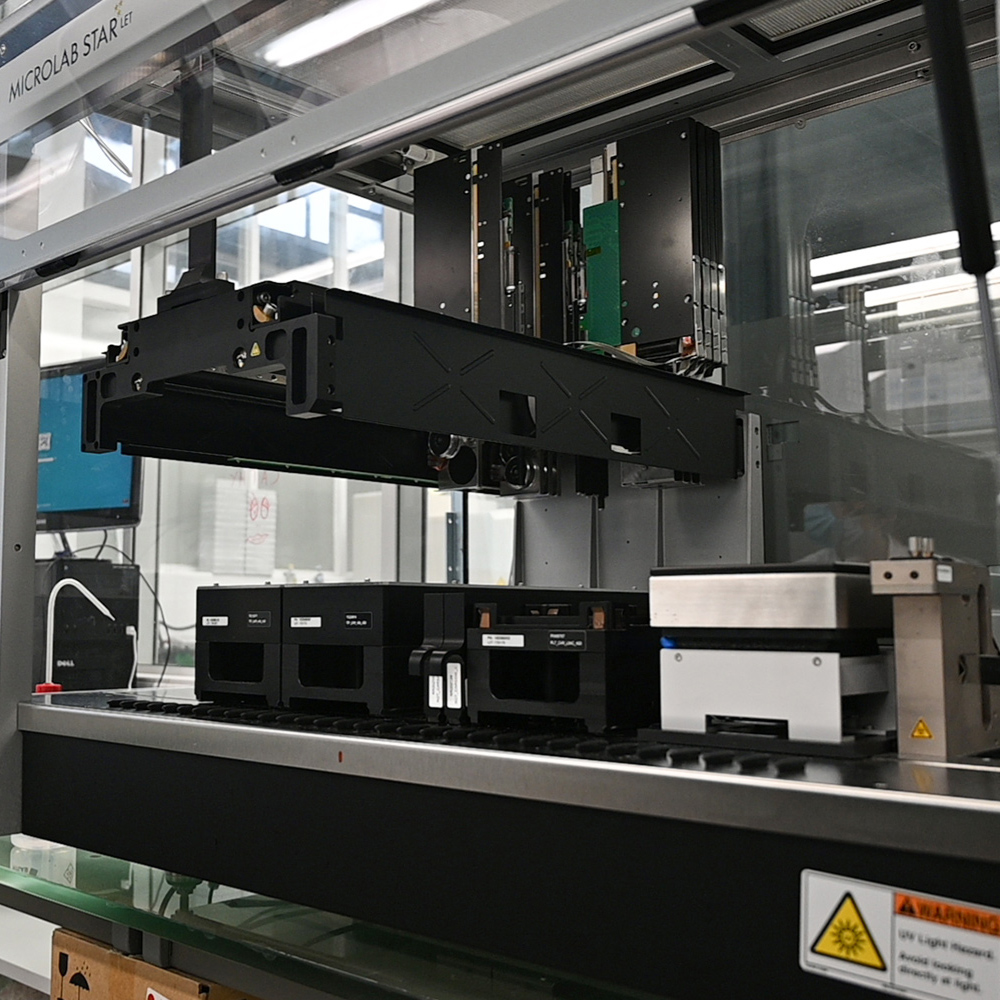
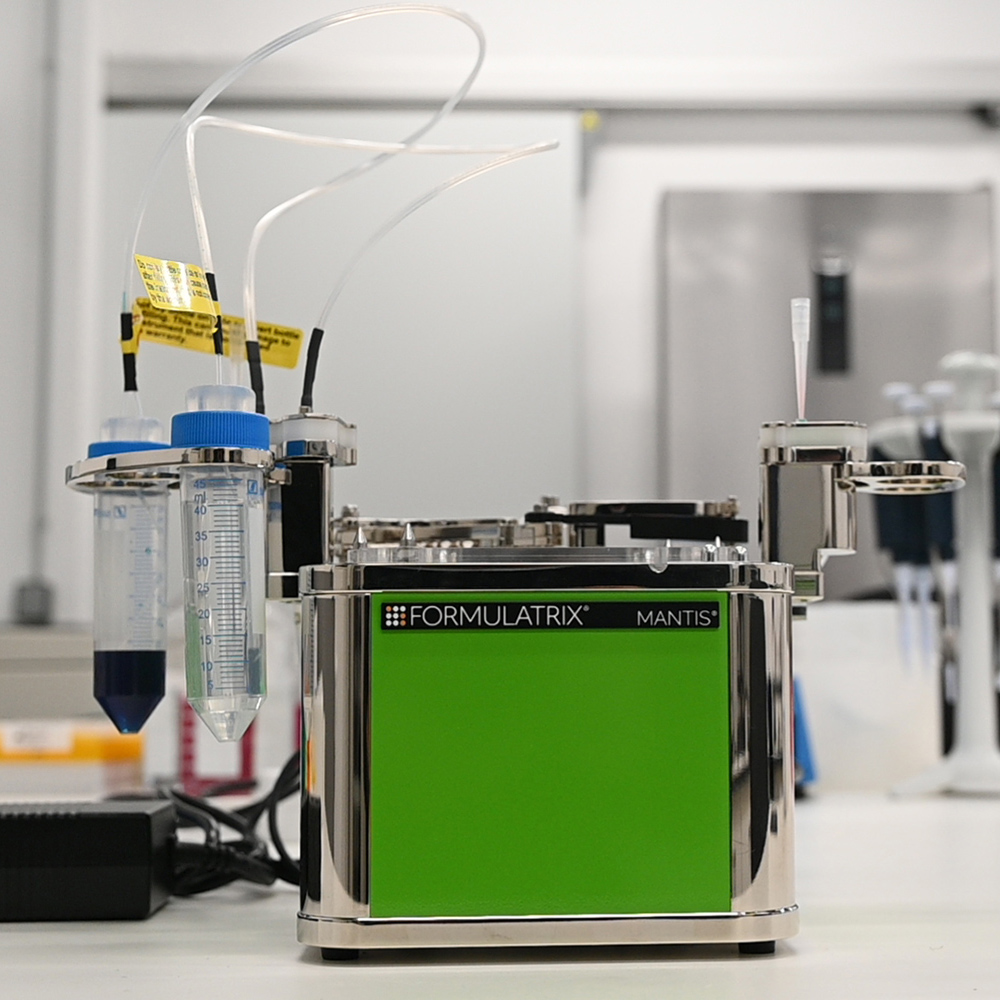
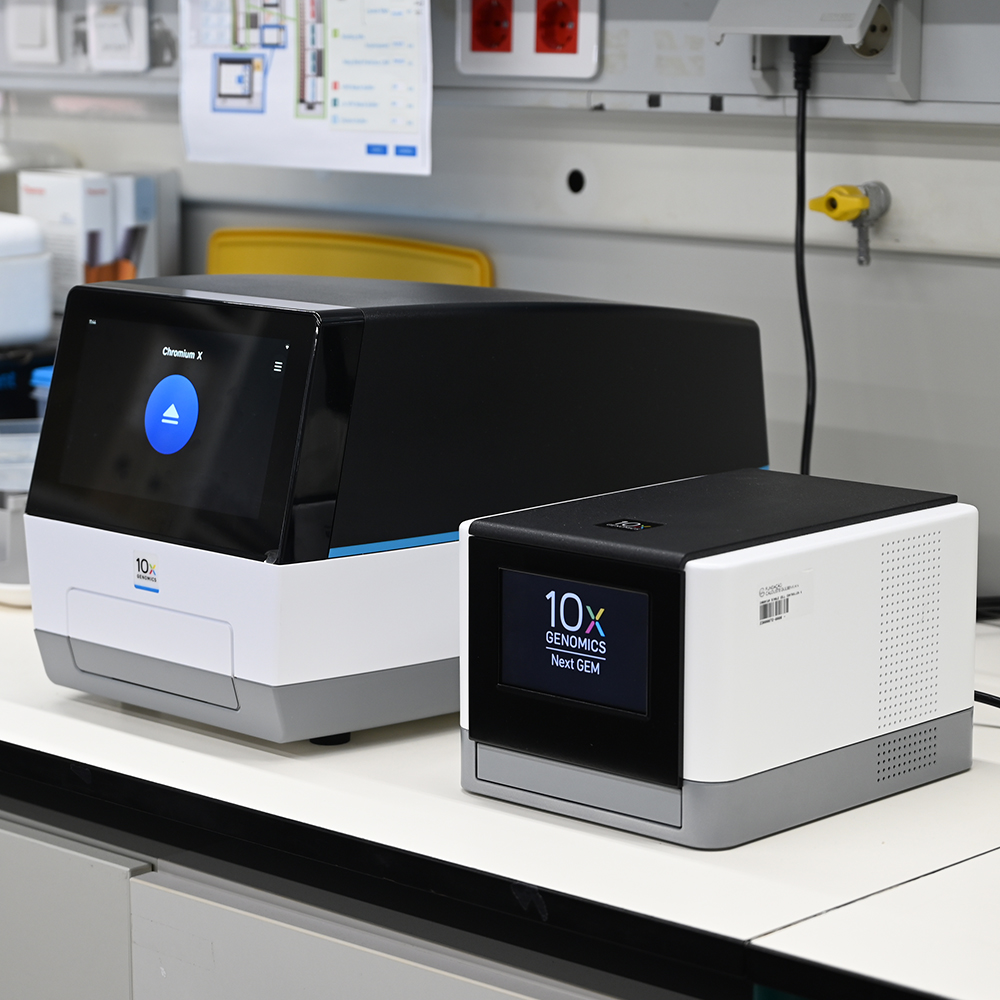
The Genomics Unit offers short read (Illumina NexSeq 2000 and Illumina MiSeq) and long read Nanopore sequencing, Library preparation, and Fragment-analyzer/TapeStation sample QC services. The unit provide a range of services: RNA-Seq, Whole Genome Sequencing, Spatial transcriptomics and Metagenomics (bacterial 16S rRNA, ITS). For Single Cell sequencing the unit operates a 10X Genomics Chromium controller and the new Chromium X capable of high-throughput experiments. We are equipped the most recent liquid handling technology (Biomek i7; Hamilton Starlet and Formulatrix Mantis).
Location: Marco Polo Wing
Contacts: +351 214407975 @: [email protected]
EQUIPMENT








SERVICES
The unit offers RNA-seq library preparation, with multiple options such as ribo-depletion (Bacterial and Eukaryotic), poly-A enrichment, 3′-Tag-Seq (QuantSeq) libraries All the libraries are constructed to retrieve the full transcript, exception for QuantSeq (3′-end sequencing). For single cell RNA-Seq please check this section
Sequencing is performed Single End (SE)100bp, PE100 (100+100) and SE150 (150+150). Optional SE50 is available under consulting.
The SMART-SEQ2 protocols we have available allow us to produce RNA-Seq libraries from very limited amounts of starting material (5ng of high-quality total RNA down to single cells). Reads cover complete transcripts and can be used to detect alternative splicing, in addition to regular differential expression.
Key features:
Requirements:
We require total RNA samples. We do not perform RNA isolations in the lab.
The unit has implemented a low-volume Nextera protocol that uses the Tn5 transposase for tagmentation-based library construction.
Key features:
Requirements:
Nanopore sequencing with Oxford Nanopore Technologies (ONT) systems enables high-throughput long-read sequencing of both DNA and RNA samples. We offer long reads lib preparation and sequencing on a MinION instrument (PromethION P2 soon), with either MinION or Flongle flow cells.
Key features:
Requirements:
The unit offers 10X Genomics Visium Spatial Transcriptomics service with the Histopathology Facility. This allows for transcriptome-wide mapping of genes expression onto tissue sections, with concomitant H&E or IF staining. The option to process both FFPE slides (human and mouse only) or Fresh frozen tissues is available. An upgrade of the technique Visium HD, bringing single-cell resolution, will be released in the coming months. Please contact us for more details.
Classical 16S V4 rRNA sequencing
This metagenomic protocol focuses on the V4 region of bacteria 16S region (primers 515F-806R) and is designed to amplify from bacteria and archaea (Walters et al., 2016). Using a 280-multiplex approach on a 2x250bp PE MiSeq run we deliver up to 50.000 reads per sample. Other primers combinations on request (e.g. V4-V5)
Key features:
Requirements:
Express Service (V3-V4) and or ITS (96 samples)
Contact us for more information.
The unit offers two possibilities: 10x genomics and one on the way Flash-seq (to be implemented).
10x Genomics is a single-cell profiling technology is becoming the most popular alternative for the analysis of large cell numbers. The platform allows for high-throughput analysis in a variety of cell types as well as single-cell nuclei. The workflow encapsulates cells or nuclei together with gel beads into nanodroplets. Different protocols are available: Gene Expression (GEX), scATAC, GEX+immune profiling and multiome (GEX + ATAC). Preliminary Single-cell data analysis is provided by Cell Ranger and Loupe Cell Browser software.
Possibilities:
For any queries please contact us.
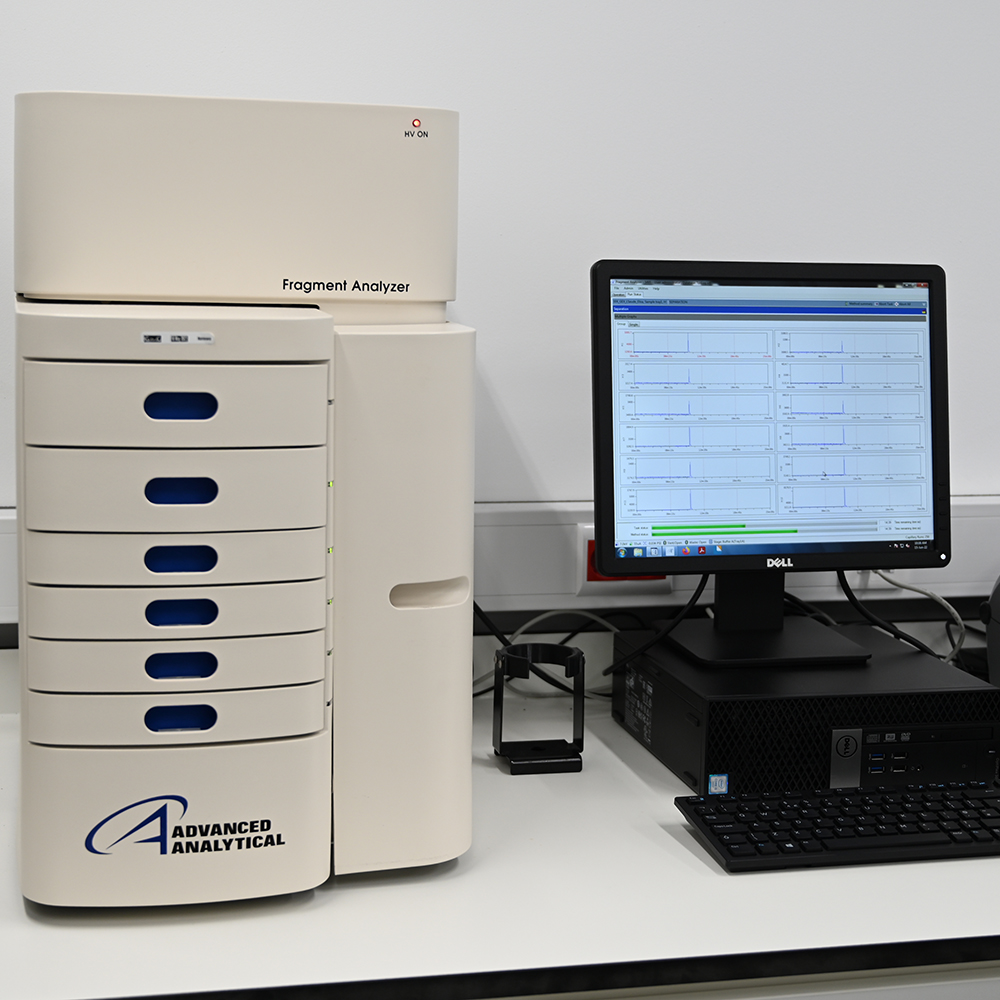
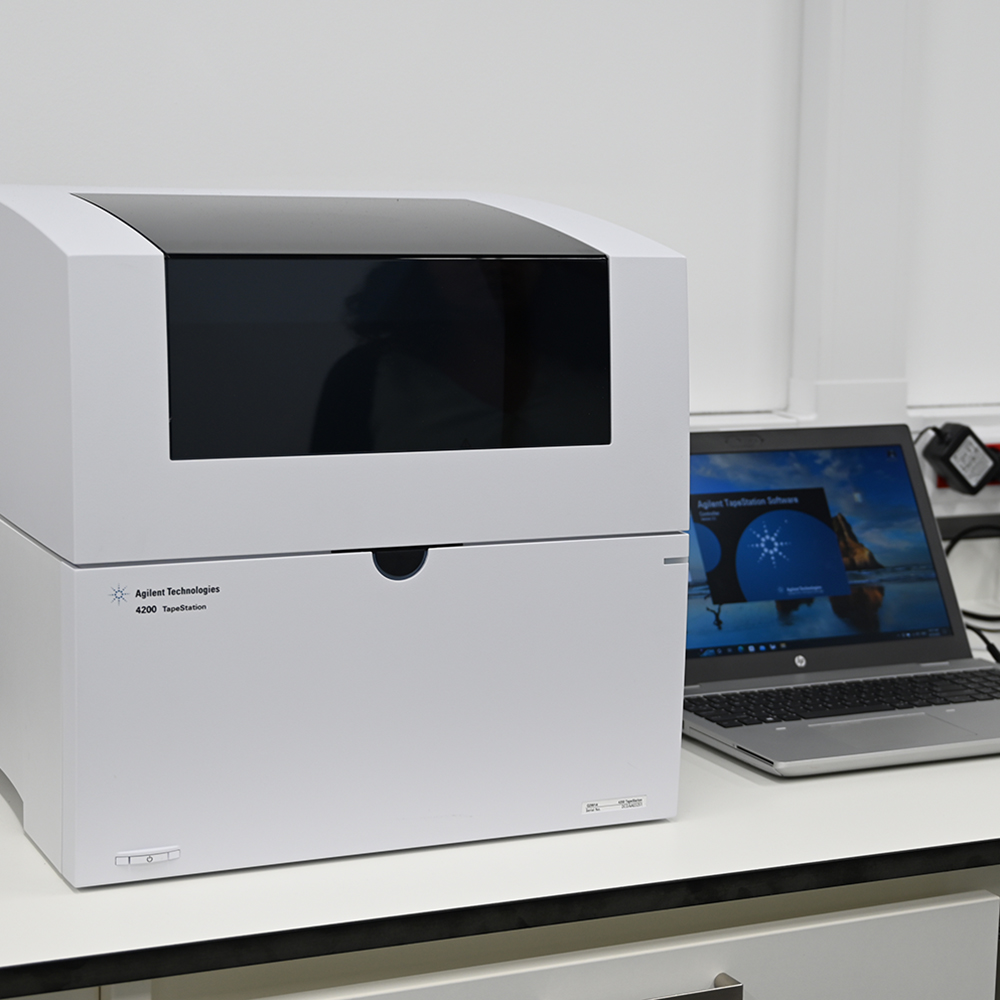
Our Nucleic Acid QC service is based on 2 instruments: Fragment analyser and TapeStation both from Agilent. Fragment analyser is more suitable for a large number of samples and TapeStation for fast QC acquisition and small number of samples ~7-8. Fragment analyser is more sensitive for low amounts of RNA/DNA while TapeStation can discriminate the sample quality in RIM (RNA) or DIM (DNA).
DNA - High Sensitivity NGS
DNA Fragment Concentration Range: 5 pg/uL - 500 pg/uL; DNA Smear Concentration Range: 50 pg/uL - 5000 pg/uL
RNA - High Sensitivity RNA
Quantitative Range (per smear) (Total RNA) 50 pg/uL - 5000 pg/uL; (mRNA) 500 pg/uL - 5000 pg/uL
TapeStation:
DNA - D1000: DNA Sizing Range: 35 bp - 1000 bp
Sample Concentration: 0.1 ng/µL - 50 ng/µL input DNA
DNA - D5000: DNA Sizing Range: 100 bp - 5000 bp
Sample Concentration: 0.1 ng/µL - 50 ng/µL input DNA
DNA - Genomic: Sizing Range: 200 bp - 60000 bp
Sample Concentration: 10 ng/µL - 100 ng/µL input DNA
RNA - High Sensitivity: Sizing Range: 100 bp - 6000 bp
Sample Concentration: 0.5 ng/µL - 10 ng/µL input RNA
PEOPLE
PUBLICATIONS
Check the papers here.